Introduction
In the previous post, we used command line tools to manipulate and study text files that contained rows and columns of data, some numeric. These kind of files are often known as CSV files–for “comma separated values”–even though the separators may be other punctuation characters, tabs or spaces. Putting data into CSV format is one way of structuring it, while still allowing it to be stored in a human- and machine-readable file. Not all data lends itself to being laid out in rows and columns, however. A different strategy for representing structure is to provide markup in the form of tags that indicate how a region of text should be displayed, what it means, or some other associated metadata. If these metadata tags are stored in the same file as the text to which they refer, they need to be syntactically distinguished from their surroundings. That is to say, it should be perfectly clear to a human or machine reader which part of the file is text and which part is tag. Here are some examples. The sentence below has two HTML (HyperText Markup Language) tags which indicate how it should be displayed in a web browser. Note the use of angle brackets and a forward slash to indicate which is the beginning tag and which is the ending one.
This is how you indicate <em>emphasis</em>, and this is how you make something <strong>stand out</strong> from its surroundings.
In XML (Extensible Markup Language), you can create tags to represent any kind of metadata you wish.
The field notes were written by <author>Frank N. Meyer</author>.
In HTML and XML, tags should be properly nested.
<outside_tag>This is <inside_tag>properly nested</inside_tag></outside_tag> <outside_tag>This is <inside_tag>not properly nested</outside_tag></inside_tag>
[UPDATE 2014. This post was changed slightly to reflect changes in WorldCat records that were made since the original post. The sense of the lesson is unchanged.]
Installation
Since markup files are plain text, we can use Linux command line tools like cat, tr, sed, awk and vi to work with them. It turns out to be sometimes difficult to match tags with regular expressions, however, so working with grep can be frustrating. We will install a special utility called xmlstarlet to alleviate some of these problems. Use man to see if you have xmlstarlet installed. If not, install it with
sudo aptitude install xmlstarlet
[UPDATE 2014. If you are using HistoryCrawler you will have to install xmlstarlet but not graphviz.]
Later in this post we are also going to visualize some relationships in the form of graphs, diagrams that show lines or arrows connecting points or labeled nodes. We will use the graphviz package for this. Use man to see if you have it installed. If not, install it with
sudo aptitude install graphviz
Getting an XML document and extracting elements
In previous posts (1, 2), we spent some time looking at the field notes written by Frank N. Meyer during a USDA-sponsored botanical expedition to South China, 1916-18. Here we are going to use the OCLC WorldCat Identities database to learn more about Meyer and the people with whom he was associated. OCLC, the Online Computer Library Center, is the organization that maintains WorldCat, a union catalog of the holdings of tens of thousands of libraries worldwide. The Identities database contains records of the 30 million persons, organizations, fictitious characters, and so on that the items in WorldCat are by or about. Start your GUI, open a terminal and start the iceweasel (or Firefox) browser in the background. The Identities page for Frank N. Meyer is at http://www.worldcat.org/identities/lccn-n83-126466. Spend some time exploring the page so you know what is on it. Now, in a new browser tab, open the XML version of the same page at http://www.worldcat.org/identities/lccn-n83-126466/identity.xml. Spend some time comparing the two pages. You want to discover how information that is presented for human consumption in the regular webpage is encoded in human- and machine-readable tags in the XML page. Note that you should be able to expand and collapse XML tags in your web browser display. In iceweasel, you do this by clicking on the little minus signs beside a particular tag. Doing this will give you a better sense of how the tags are nested. Now that we have a sense of the structure of the XML document, we can try extracting some of this information using command line tools. In the terminal, use wget to download a local copy of the XML file, then use the xmlstarlet el command to get a listing of the elements in the file.
wget http://www.worldcat.org/identities/lccn-n83-126466/identity.xml xmlstarlet el identity.xml | less
Note that for associated names, we see the following pattern repeated:
Identity/associatedNames/name Identity/associatedNames/name/normName Identity/associatedNames/name/rawName Identity/associatedNames/name/rawName/suba Identity/associatedNames/name/rawName/subb Identity/associatedNames/name/rawName/subd
Each of these lines represents a ‘path’ to a particular element in the XML document. Looking at the XML display in iceweasel we can see how associated names are tagged. We see that the normName field contains the LCCN. This is the Library of Congress Control Number, a unique identifier. Frank N. Meyer’s LCCN is n83126466. The human-readable name is stored in the rawName/suba field, with optional information in rawName/subb. Dates are in rawName/subd.
Selecting information from an XML document
We can pull information out of an XML file using the xmlstarlet sel command. For example, if we wanted to count the number of associated names, we would type the following. The -t option tells xmlstarlet to return plain text; the -v (value) option tells it what we are looking for.
xmlstarlet sel -t -v "count(/Identity/associatedNames/name)" -n identity.xml
Here we are more interested in using xmlstarlet to parse the XML file, that is, to find information and extract it. As a first attempt, we try matching (-m) the associated names and pulling out the values (-v) of the normName fields. The -n option puts newlines where we want them.
xmlstarlet sel -t -m "/Identity/associatedNames/name" -v "normName" -n identity.xml
The output that we get looks like the following.
lccn-n79008243 lccn-n83126465 lccn-n80145310 nc-united states$government printing office lccn-n85140459 lccn-n85335475 lccn-n50015525 np-kelsey, harlan p lccn-n79076676 lccn-n85800990
While we are at it, we can also grab information from the rawName fields. We modify our command to do that, outputting the results in a colon-separated table. The -T option says we want to output plain text. The -o option provides our output separators. Note that we are also including escaped quotation marks around our name fields. This will help us later when we further manipulate the information we are extracting.
xmlstarlet sel -T -t -m "/Identity/associatedNames/name" -v "normName" -o ":\"" -v "rawName/suba" -o " " -v "rawName/subb" -o "\"" -n identity.xml
Our output now looks like this:
lccn-n79008243:"United States Dept. of Agriculture" lccn-n83126465:"Cunningham, Isabel Shipley " lccn-n80145310:"United States Bureau of Plant Industry" ... lccn-n85800990:"Smith, Erwin F. "
The blank spaces in the rawName fields will cause us problems later, so we are going to use tr to eliminate those. We will use grep to get rid of the entries that don’t have proper LCCNs. Finally, we will package everything up into a convenient Bash shell script. Use a text editor to create the following file, and name it get-assoc-names.sh.
#! /bin/bash xmlstarlet sel -T -t -m "/Identity/associatedNames/name" -v "normName" -o ":\"" -v "rawName/suba" -o " " -v "rawName/subb" -o "\"" -n $1 | grep 'lccn' | tr -d ' '
Now you can change the permissions and try executing the script as follows. The last command shows how you can use cut to pull out just the names.
chmod 744 get-assoc-names.sh ./get-assoc-names.sh identity.xml ./get-assoc-names.sh identity.xml | cut -d':' -f2
We can also write a small script to pull out the LCCN and rawName for the identity that the file is about (in this case, Frank N. Meyer). Look at the XML display in your browser again. In this case, we have to use the ‘@’ character to specify the value for a tag attribute. Use a text editor to write the following script, save it as get-name.sh, change the file permissions and try executing it.
#! /bin/bash xmlstarlet sel -T -t -v "/Identity/pnkey" -o ":\"" -v "/Identity/nameInfo[@type='personal']/rawName/suba" -o "\"" -n $1 | tr -d ' '
Plotting a directed graph
If we want to visualize the relationships between a set of entities, one way is to create a graphical figure that shows the entities as dots or nodes, and the relationships as lines or arrows. In mathematics, this kind of figure is known as a graph (not to be confused with the other sense of the word, which usually refers to the plot of a function’s output). If the connection between two entities is directional (an arrow, rather than a line), the graph is called a digraph, or directed graph. Suppose that John Doe has some kind of relationship to Jane Doe: he might be her son, nephew, husband, uncle, Facebook friend, whatever. If we want to visualize this relationship with the Graphviz software package, we start by creating a file that looks like the following. Use a text editor to create the file and save it as example.dot.
digraph G {
"JohnDoe" -> "JaneDoe";
}
Next we use the Graphviz neato command to convert the description of the digraph into a picture of it, and save the output as a .PNG graphics file. Finally we use the display command to show the picture (and put the process in the background using an ampersand).
neato -Tpng -Goverlap=false example.dot > example.png display example.png &
The resulting image looks like this:  In order to lay out the graph, neato uses what is called a ‘spring model’. Imagine all of the nodes of the graphs are weights, and all of the arrows connecting them are compression springs that are trying to push the weights apart. By simulating this process, neato arrives at a figure where the nodes are separated enough to read them, but not so far as to waste space. Now suppose we want to graphically represent the relationship between the main identity in our XML file (i.e., Frank N. Meyer) and all of the identities that he is associated with. We can use a Bash script to build the digraph file automatically from the identity.xml file. We will do this in stages. We start by using the echo command to print out all of the lines of our file. Note we have to use the escape character to include one set of quotation marks inside of another. Use a text editor to create the following file and name it build-digraph.sh
In order to lay out the graph, neato uses what is called a ‘spring model’. Imagine all of the nodes of the graphs are weights, and all of the arrows connecting them are compression springs that are trying to push the weights apart. By simulating this process, neato arrives at a figure where the nodes are separated enough to read them, but not so far as to waste space. Now suppose we want to graphically represent the relationship between the main identity in our XML file (i.e., Frank N. Meyer) and all of the identities that he is associated with. We can use a Bash script to build the digraph file automatically from the identity.xml file. We will do this in stages. We start by using the echo command to print out all of the lines of our file. Note we have to use the escape character to include one set of quotation marks inside of another. Use a text editor to create the following file and name it build-digraph.sh
#! /bin/bash
echo "digraph G {"
echo " \"John Doe\" -> \"Jane Doe\";"
echo "}"
Change the permissions and try executing your shell script with the following commands.
chmod 744 build-digraph.sh ./build-digraph.sh
Instead of echoing the statements inside our digraph, however, we want to construct them using the xmlstarlet bash scripts that we just made. First we input the identity file on the command line and grab the name from it. Use a text editor to edit build-digraph.sh so it now looks as follows.
#! /bin/bash
NAME=$(./get-name.sh $1 | cut -d':' -f2)
echo "digraph G {"
echo " "${NAME}" -> foobar;"
echo "}"
Try running it with
./build-digraph.sh identity.xml
Now we want to create one line in our digraph file for each associated name. This is clearly a job for the for loop. Use a text editor to edit build-digraph.sh so it now looks as follows.
#! /bin/bash
NAME=$(./get-name.sh $1 | cut -d':' -f2)
echo "digraph G {"
for ANAME in $(./get-assoc-names.sh $1 | cut -d':' -f2)
do
echo " "${NAME}" -> "${ANAME}";"
done
echo "}"
To see if it works as expected, try running it with
./build-digraph.sh identity.xml
Your output should look like this:
digraph G {
"Meyer,FrankNicholas" -> "UnitedStatesDepartmentofAgriculture";
"Meyer,FrankNicholas" -> "Cunningham,IsabelShipley";
"Meyer,FrankNicholas" -> "UnitedStatesBureauofPlantIndustry";
"Meyer,FrankNicholas" -> "UnitedStatesOfficeofForeignSeedandPlantIntroduction";
"Meyer,FrankNicholas" -> "Fairchild,David";
"Meyer,FrankNicholas" -> "Wilson,ErnestHenry";
"Meyer,FrankNicholas" -> "PopulationReferenceBureau";
"Meyer,FrankNicholas" -> "Smith,ErwinF.";
}
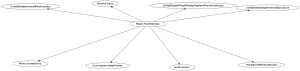
That looks good, so we send the output to a .dot file, run it through Graphviz neato and display.
./build-digraph.sh identity.xml > anames.dot neato -Tpng -Goverlap=false anames.dot > anames.png display anames.png &
If all went well, the output should look like this:
Trees are graphs, too
Recall that XML files are structured so that tags are properly nested inside one another. We can visualize this containment relationship as a digraph, where we have arrows from outside_tag to inside_tag. In the next section we will use xmlstarlet to extract the structure of our XML file, then Graphviz to plot it in the form of a digraph. Instead of using neato, we will use a different Graphviz plotting routine, dot, which is more appropriate for tree-like figures. Using the -u option, we can eliminate redundant tags when we pull the elements out of an XML file with xmlstarlet el.
xmlstarlet el -u identity.xml | less
Look at the previous listing. In order to convert it into the proper form for Graphviz, we need to turn the forward slashes into arrows. We have a more tricky problem, however. We need to lose everything in a given line except the last two tags and the slash between them. Think about this until you understand why it is the case. We will use grep to pull out the last two tags, separated by a slash. In order to match one or more copies of something that is not a slash, we use the following pattern
([^/]+)
So we want a string of characters that are not a slash, followed by a slash, followed by another string of characters that are not a slash, followed by the end of the line. And we want to match only that (the -o option for extended grep). The following pipeline does what we want.
xmlstarlet el -u identity.xml | grep -E -o '([^/]+)/([^/]+)$' | less
Now we use sed to replace each slash with the correct characters for Graphviz. Our XML tags contain some colons that will be confusing to Graphviz if we leave them in the tags. We are going to translate these into underscores for the purpose of making graph labels. Try the following version of the pipeline.
xmlstarlet el -u identity.xml | grep -E -o '([^/]+)/([^/]+)$' | sed 's/\//->/g' | tr ':' '_' | less
The output looks good, so we can bundle everything into a bash script called build-xml-tree.sh
#! /bin/bash
echo "digraph G {"
for LINK in $(xmlstarlet el -u $1 | grep -E -o '([^/]+)/([^/]+)$' | sed 's/\//->/g' | tr ':' '_')
do
echo ${LINK}";"
done
echo "}"
Try running the shell script so you can make sure the output looks right.
chmod 744 build-xml-tree.sh ./build-xml-tree.sh identity.xml | less
Finally we lay out our digraph with dot. The -Grankdir option tells Graphviz that we want to use a left-right layout rather than a top-down one. This will give us a figure that is more easily compared with our web browser display.
./build-xml-tree.sh identity.xml > xmltree.dot dot -Tpng -Goverlap=false -Grankdir=LR xmltree.dot > xmltree.png display xmltree.png &
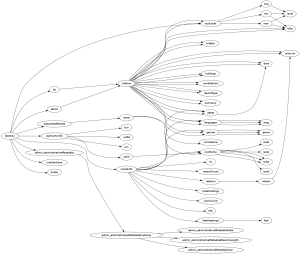
The resulting digraph looks like this.
Study this figure for a few minutes. Because it encodes information about the XML file in a different way than the XML display in the web browser does, it makes it easier to see some things. For example, it is obvious when you look at the xmltree.png figure that some tags, like oclcnum or citation, may be children of more than one parent. What else can you discover about the XML file by studying the graph visualization of it?


0 Comments